
HTRF EPIgeneous Human and Mouse Total Histone H4 Detection Kit, 500 Assay Points
This HTRF kit allows the cell-based quantitative detection of Total Histone H4.
| Feature | Specification |
|---|---|
| Application | 細胞シグナル伝達 |
| Sample Volume | 10 µL |
This HTRF kit allows the cell-based quantitative detection of Total Histone H4.


Loading...
Product information
Overview
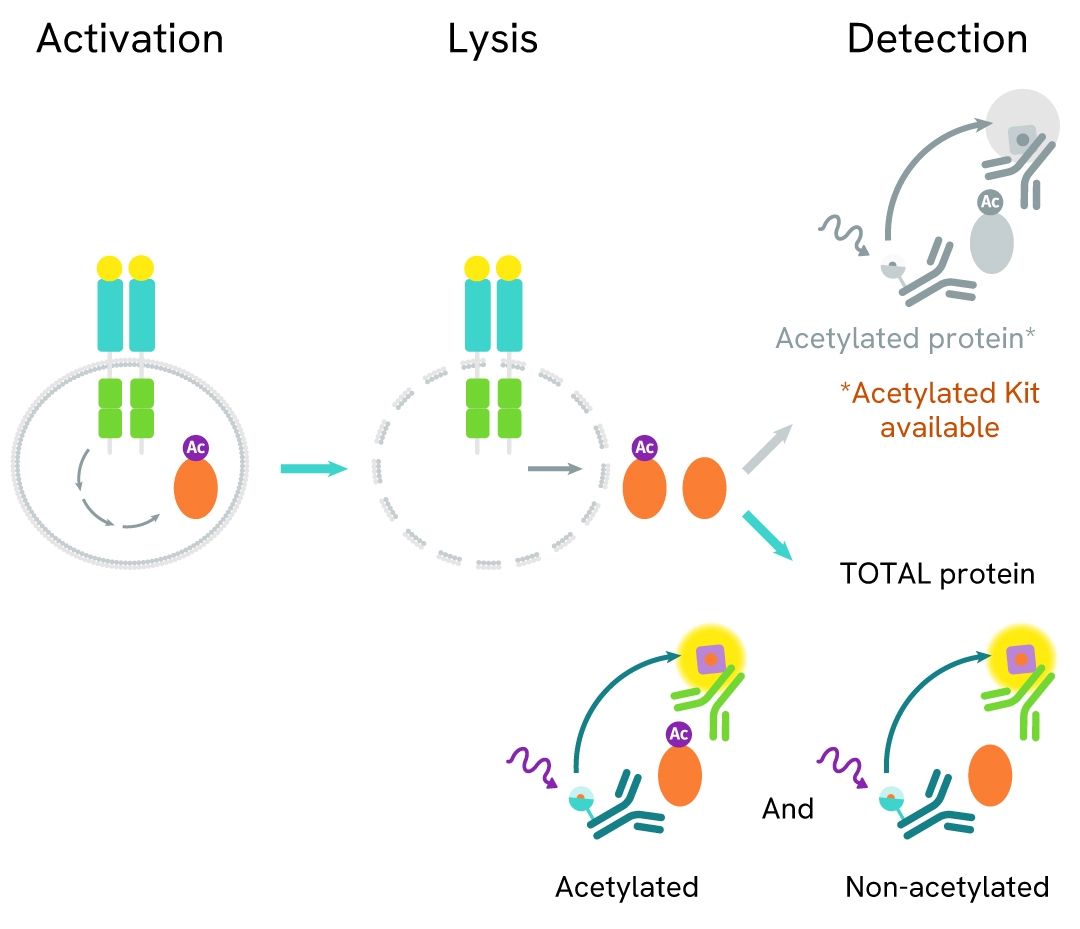
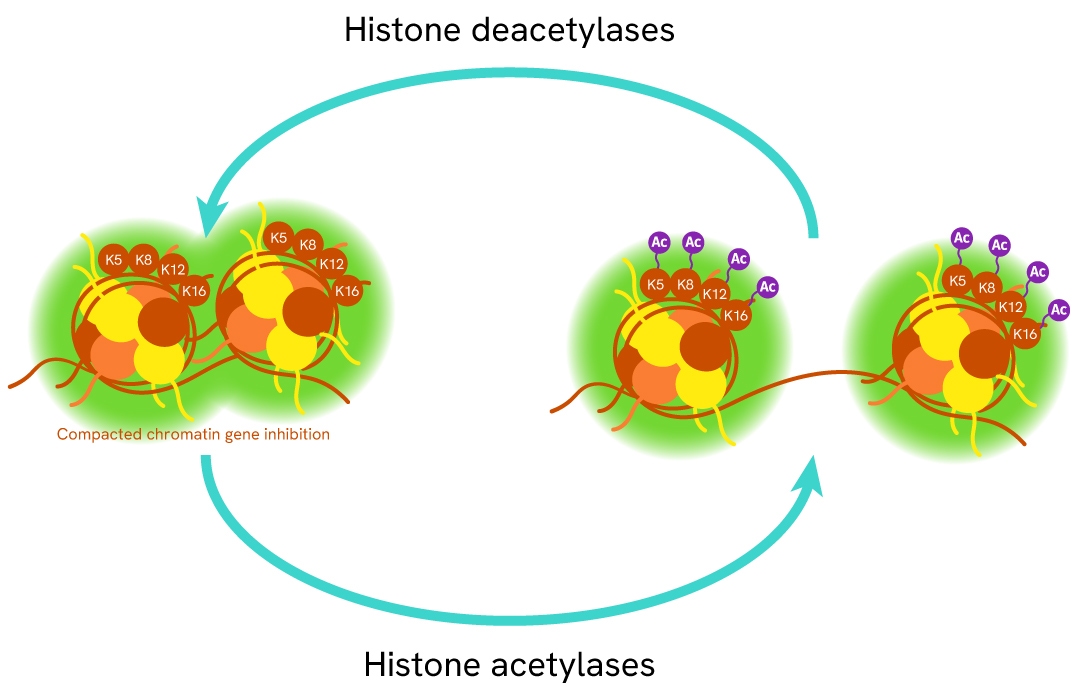
Histone 4 is one of the core histone proteins involved - along with other histones - in the packaging of DNA into a condensed chromatin state. Histone H4 function is dependent on its acetylation state (1 acetyl group per lysine) at several sites (Lys5, Lys8, Lys12, and Lys16) on the N-terminal tail. The mis-regulation of histone acetylation is closely associated with cancer initiation and progression, making histone acetylase and deacetylase enzymes promising drug targets.
HTRF assays offer many advantages over other technologies:
- Homogeneous add-and-read format
- No wash steps
- Low background
- Straightforward miniaturization from 96- or 384-well microplates to high density assay formats such as 384-well low volume and 1536-well plates
- Stable signal, providing flexibility in readout time or size of assays
How it works
Total Histone H4 assay principle
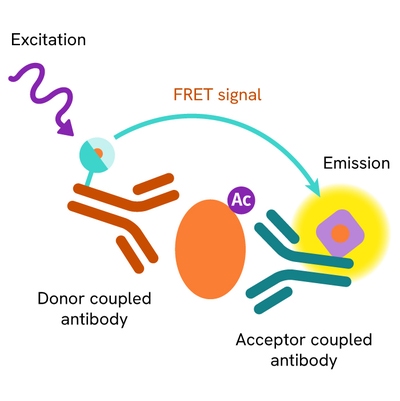
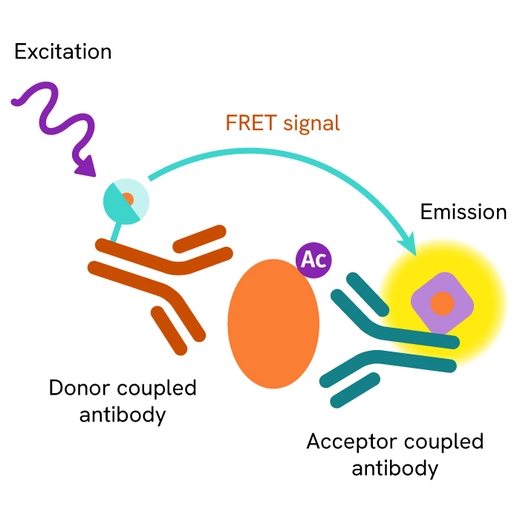
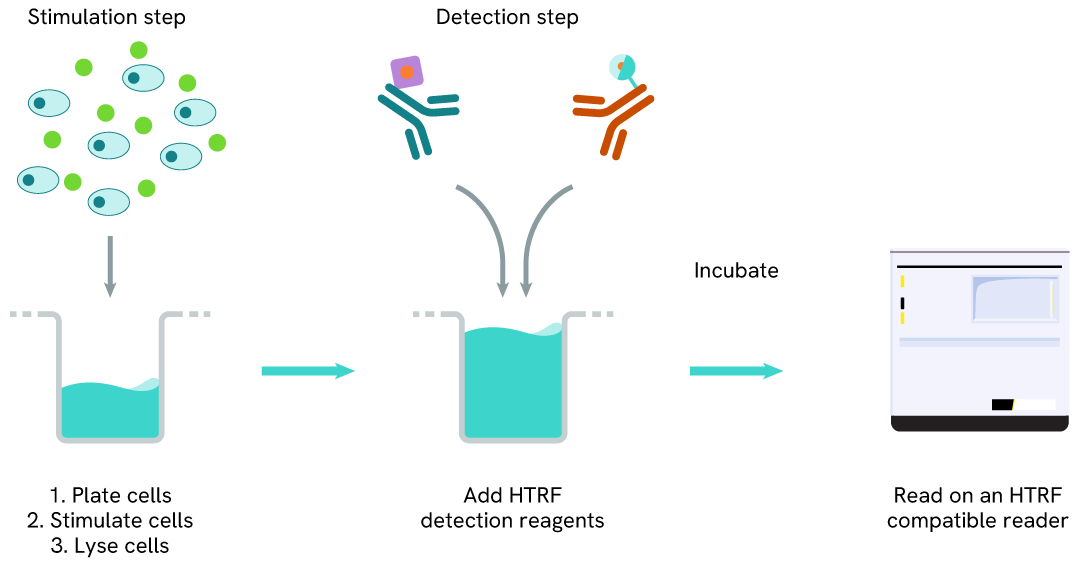
The Total Histone H4 assay measures Histone H4 levels in cells. Unlike Western Blot, the assay is entirely plate-based and does not require gels, electrophoresis, or transfer. The assay uses 2 antibodies, one labeled with a donor fluorophore and the other with an acceptor. Both antibodies are highly specific for a distinct epitope on the protein.
In the presence of Histone H4, this enables an immune-complex formation involving both labeled antibodies, and which brings the donor fluorophore into close proximity to the acceptor, thereby generating a FRET signal. Its intensity is directly proportional to the concentration of total protein present in the sample.

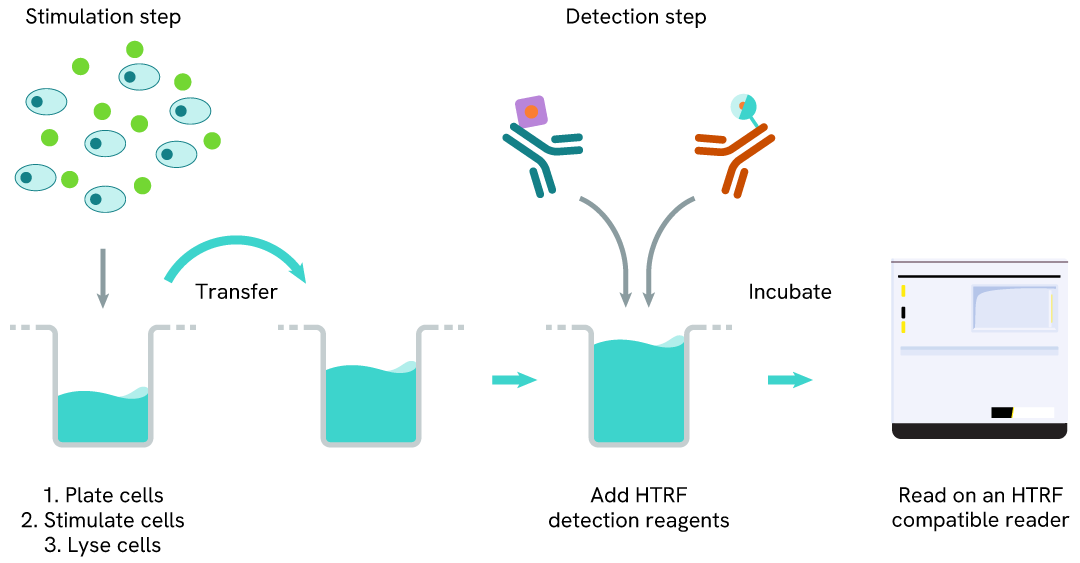
Total Histone H4 two-plate assay protocol
The two-plate protocol involves culturing cells in a 96-well plate before lysis, then transferring lysates into a 384-well low volume detection plate before the addition of Total Histone H4 HTRF detection reagents. This protocol allows the cells' viability and confluence to be monitored.
Total Histone H4 one-plate assay protocol
Detection of Total Histone H4 with HTRF reagents can be performed in a single plate used for culturing, stimulation, and lysis. No washing steps are required. This HTS designed protocol allows miniaturization while maintaining robust HTRF quality.
Assay validation
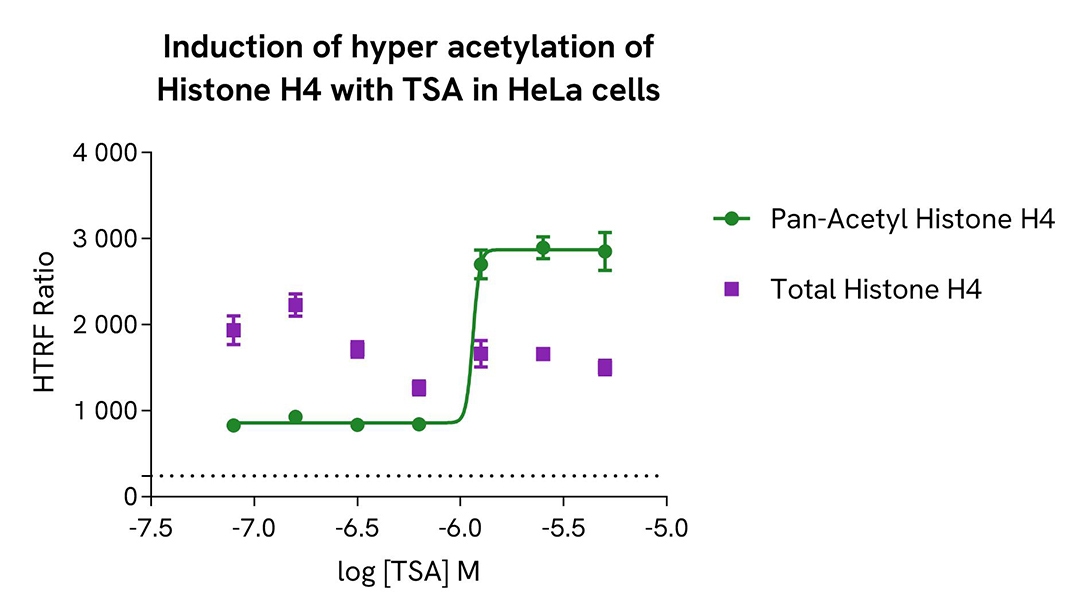
Induction of Histone H4 hyper acetylation with Trichostatin A in HeLa cells
HeLa cells were seeded in a 96-well culture-treated plate (12,500 cells/well) in complete culture medium, and incubated overnight at 37°C, 5% CO2. The cells were treated for 18h with increasing concentrations of the histone deacetylase inhibitor Trichostatin A (TSA), in order to increase acetylated Histone H4 levels in cells.
After treatment, the cells were lysed with 160 μL of lysis buffer A-part 1 and incubated for 45 minutes at RT under gentle shaking. After incubation, 40 μL of Lysis Buffer A-part 2 were added. For the detection step, 10 µL of cell lysate (diluted at 1/4) were transferred into a 384-well low volume white microplate and 10 µL the HTRF Total Histone H4 or HTRF Pan-Acetyl Histone H4 (Revvity, #62H4PANACPAE/PAD/TDA) detection reagents were added. After a 2h-incubation, the HTRF signal was recorded with an Envision Nexus reader.
In parallel, cell viability was assessed with the ATPlite 1step Kit (Revvity, #6016736/1/9). 5 µL of the same cell lysate were transferred into an HTRF 96-well low volume white plate, and 25 µL of ATPlite 1step detection reagent. After a 10-min incubation, the luminescence signal was measured with an Envision Nexus reader.
As expected, TSA promoted the hyper acetylation of Histone H4, without any significant effect on the expression level of the total Histone H4 protein. In addition, TSA did not induce cytotoxic effects, as measured by the cell viability indicator ATPLite (data not shown).
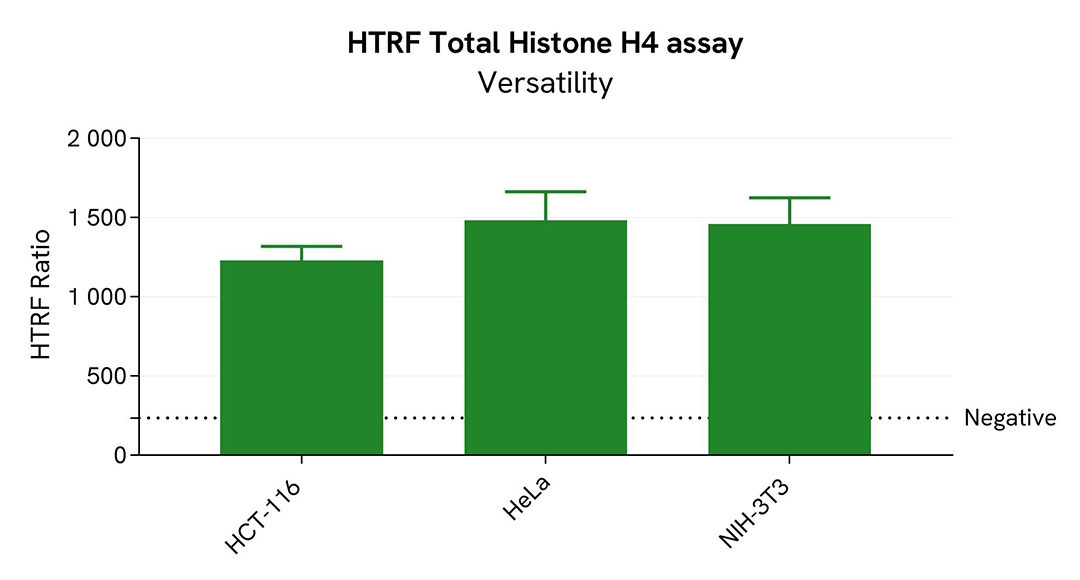
Assessment of Histone H4 hyper acetylation in various cell lines
Two human cell lines, HeLa and HCT-116, and one mouse cell line, NIH-3T3, were seeded in a 96-well culture-treated plate (12,500 cells/well) in complete culture medium, and incubated overnight at 37°C, 5% CO2. The cells were treated for 18h with Trichostatin A (TSA) (1.25 µM for HeLa and NIH-3T3, and 1 µM for HCT-116), a histone deacetylase inhibitor that promotes higher histone acetylation levels.
After treatment, the cells were lysed with 160 μL of lysis buffer A-part 1 and incubated for 45 minutes at RT under shaking. After incubation, 40 μL of Lysis Buffer A-part 2 were added. For the detection step, 10 µL of cell lysate (diluted at 1/8) were transferred into a 384-well low volume white microplate and 10 µL of the HTRF Total Histone H4 detection reagents were added. After a 2h-incubation, the HTRF signal was recorded with an Envision Nexus reader.
The HTRF Pan-Acetyl Histone H4 assay efficiently detected Histone H4 in various cellular models expressing different levels of the protein.
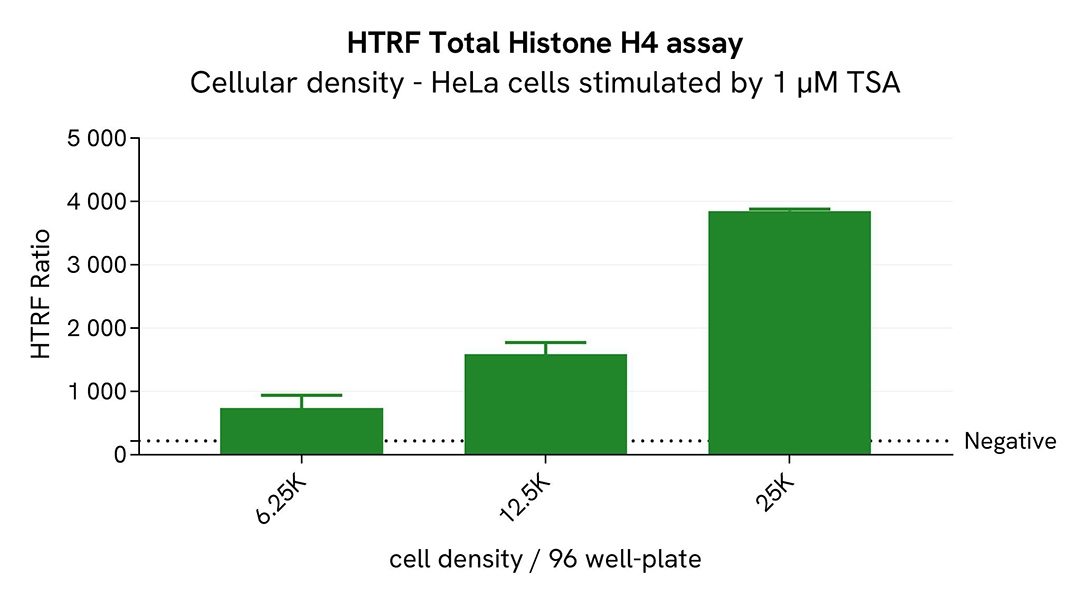
Broad dynamic range detection of Total Histone H4 assay
Various densities of HeLa cells were seeded in a 96-well culture-treated plate in complete culture medium, and incubated overnight at 37°C, 5% CO2. The cells were treated for 18h with increasing concentrations of the histone deacetylase inhibitor Trichostatin A (TSA), in order to increase acetylated Histone H4 levels in cells.
After treatment, the cells were lysed with 160 μL of lysis buffer A-part 1 and incubated for 45 minutes at room temperature under shaking. After incubation, 40 μL of Lysis Buffer A-part 2 were added. For the detection step, 10 µL of cell lysate (diluted at 1/4) were transferred into a 384-well low volume white microplate and 10 µL of the HTRF Pan-Acetyl Histone H4 detection reagents were added. After a 2h-incubation, the HTRF signal was recorded with an Envision Nexus reader.
The results show a successful detection of Pan-Acetyl Histone H4 over a broad range of cellular densities.
Simplified pathway
Histone H4 roles in DNA replication
Histones are a family of proteins with a key role in the organization and regulation of DNA. Five types exist (H1, H2A, H2B, H3, and H4) and they act as spools for DNA to wrap around in a condensed chromatin state. Individually, histones are rather small (>20 kDa, except for H1 which is 31-33 kDa), but they assemble in larger octameric nucleosome particles made of two copies of all histone types except H1. Each nucleosome carries an average of two turns of condensed DNA which sticks to the structure. This is due to the overall positive charges carried by the histones because of their positively-charged amino acids. Histones can be subject to modifications such as methylation, acetylation, phosphorylation, and ubiquitination, which all affect their ability to assemble, disassemble, and stick to DNA. All these properties regulate the physical access of proteins to the genome. Because of the way histones condense and wrap DNA around themselves, they oversee which portions of the genome are accessible to transcription enzymes and other partners that can modify or replicate it. By sliding along the DNA sequence, histones cover and uncover different sections, thus prompting the expression of new sequences and the repression of others. For these reasons, histone modifications are critical to gene silencing and expression.
This role of histones in DNA regulation is postulated under the “Histone code” hypothesis and is being increasingly studied in the context of diseases like cancer, where gene mutation and/or silencing is key. H3 and H4 acetylations are the most investigated histone modifications, and have been found to promote the transcription of genes consistently.
Specifications
| Application |
Cell Signaling
|
|---|---|
| Brand |
HTRF
|
| Detection Modality |
HTRF
|
| Lysis Buffer Compatibility |
EPIgeneous Lysis Buffer A
|
| Molecular Modification |
Total
|
| Product Group |
Kit
|
| Sample Volume |
10 µL
|
| Shipping Conditions |
Shipped in Dry Ice
|
| Target |
Histone H4
|
| Target Class |
Epigenetics
|
| Target Species |
Human
Mouse
|
| Technology |
TR-FRET
|
| Therapeutic Area |
Immuno-oncology
Oncology
|
| Unit Size |
500 assay points
|
Resources
Are you looking for resources, click on the resource type to explore further.
Loading...


How can we help you?
We are here to answer your questions.






























